
Case study: Automating large scale biomedical simulation
This case study describes my contribution within the context of the project's relevant results. My work for @neurIST took part mostly during my time at NEC and was finished at CMM as subcontract work for the project.
Challenges for automating biomedical toolchains
@neurIST supports three types of biomechanical characterisations: shape analysis, blood flow analysis, and wall stress analysis. The @neurIST biomedical toolchain starts with raw medical images, and converts them into a clean surface of the vasculature around the aneurysms. Then, for instance for blood flow analysis, it defines appropriate boundary conditions, generates input for a CFD simulation package, runs the simulation, and extracts the needed characterisations from the solution.
It was clear very soon that a high degree of automation of this toolchain was needed to cut down processing time to a feasible limit. Automation also reduces differences in results obtained by different people ("operators"), which is very important for the statistical analysis of these results. The primary challenges for automation are rooted in
- inappropriate data quality, e.g. low image resolution or contrast
- heterogeneity of data, e.g. using different protocols for medical imaging
- lack of simple means to translate intuitive concepts into clear algorithmic specifications, e.g. defining the neck of an aneurysm as the "natural" separation between aneurysm and healthy parent vessel.
Limited resolution or contrast of images may give rise to topologically incorrect geometric representations, such as almost touching vessels leading to a erroneous connection in the geometric model. Heterogeneous image acquisition protocols are a challenge for automatic segmentation. Missing formalisation of intuitive geometric concepts make it difficult for a computational tool to offer high-level methods (like define the neck of the aneurysm or clip the geometry at 10 vessel diameters from the aneurysms) and force the user to tediously combine many low-level operations to yield the desired effect.
Automation, semi-automation, ...
Tackling these challenges, the project succeed to automate a considerable number of steps. In addition, where some step could not (yet?) be fully automated, the necessary manual work could often be semi-automated by providing appropriate tools.
Some key examples of automation or semi-automation steps that @neurIST implemented are
- Using an advanced segmentation method (GAR) that could be adapted to the diverse protocols
- Providing tools to correct erroneously segmented parts of the geometry (like touching vessels erroneously connected)
- Provide a means to control the geometry, set clipping planes and link to boundary conditions, by using skeletons of the vasculature
- Generate the input for the simulations from a common high-level description of the computational problem (denoted by abstract problem description or APD, see below.)
To give an overall idea on the level of automation achieved: To date, over 260 clinicians used this toolchain successfully to process a (simple) case; under guidance of an expert, but essentially on their own.
... and human training
This level of automation helped a lot, yet it was not sufficient to guarantee comparable results across a dozen different people involved with the actual case processing. In order to reach this goal, the simulation team wrote a very detailed manual, organised several hands-on training workshop, and frequently compared results. The comparison of results was done with particular care on a set of 6 so-called demonstrator cases, where the deviations between different operators where investigated for each biomechanical characterisation value. These trainings and comparisons took place in several iterations until the results were consistent enough.
Multi-case processing
Multi-case analyses are the real proof of automation capabilities: When trying to process hundreds of cases at once, every tiny manual mouse click still necessary for a single case run hurts painfully. With abstract problems descriptions, the project has developed a means to store high-level descriptions of computational problems in a way that enables their automatic manipulation and processing.
My role in automating the biomedical simulation toolchain
Concepts
I developed general concepts for enabling automation, closely linked to the specification of processes and data entities. In particular, I worked out in detail which information was needed by each process to produce its output, which processes could be automated completely, and how the input data needed to be organised to do so. My work was guided by the maxim Generate what can be generated, take great care of the rest.
As pivotal element of automation, I introduced the concept of an abstract problem description (APD). This is a high-level, application-neutral description of a computational problem (in our domain aneurysm bio-mechanics). I defined an XML representation of APDs and a library to manage them.
From abstract to concrete
Despite its abstractness, an APD contains sufficient information to automatically generate input for concrete applications. I implemented such bespoke generators (or preprocessors) transforming APDs into input for simulation packages.
For blood flow analysis, these are
- A preprocessor for ANSYS CFX, generating CFX and ANSYS ICEM command files to automate 3D mesh generation and simulation, in collaboration with Justin Penrose (ANSYS Europe Ltd.) and Alberto Marzo (University of Sheffield)
- A preprocessor for the DC Lattice-Boltzmann (LB) solver, generating a 3D LB voxel mesh with appropriately labelled regions in collaboration with Jörg Bernsdorf, Dinan Wang and Carsten Neff (NEC Europe Ltd.)
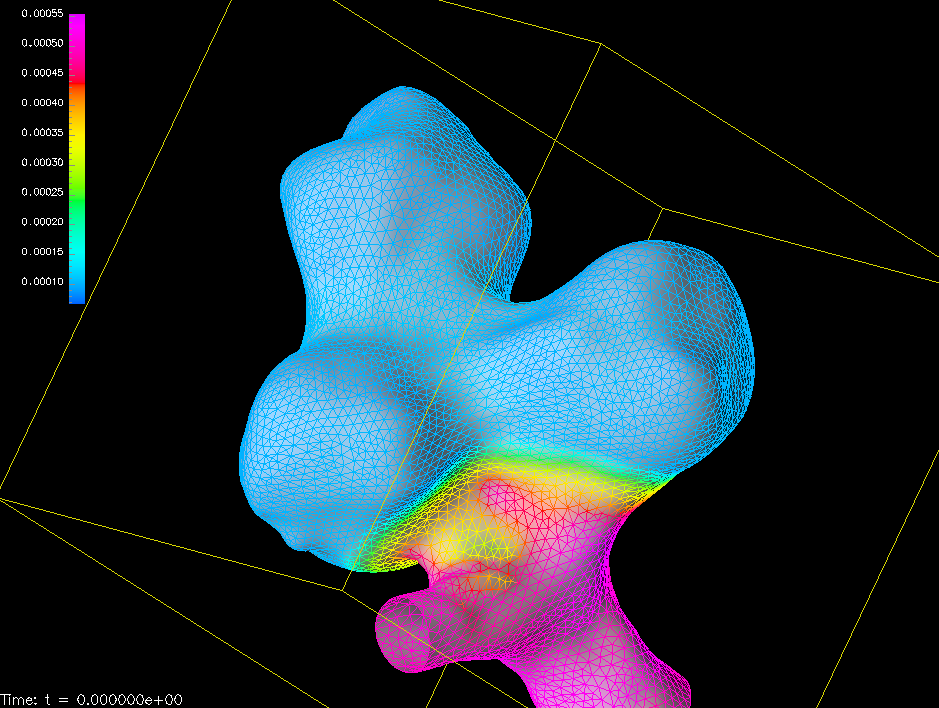
For the wall mechanics analysis, I developed similar preprocessors, supporting Jens Georg Schmidt's membrane code FEANOR and ANSYS Mechanical. The aneurysm wall is normally too thin to be measured by medical imaging, and is set to predefined value (much thinner than the healthy vessel wall). Using the normal thickness value for the vessel wall stabilises the numerical solution process. Hence, the preprocessors generate a transition region for a smooth transition from vessel wall to aneurysm wall thickness.
Algorithmic and software developments
These preprocessors feature some useful algorithmic developments, for instance a clean clipping, localised to the current vasculature branch using topological neighbourhood (in contrast, a simple geometric clipping at a cutting plane could result in cutting off unrelated parts).
I developed a number of mesh manipulation and mesh checking plug-ins (DLLs) for the graphical frontend @neuFuse (developed by B3C), which is the central tool for working with the biomechanical toolchain.
The software was implemented using C++ using generic programming, Thus, I could reuse the components of the mesh algorithms library GrAL. By employing a simple GrAL-conforming wrapper around the VTK data structures used in the @neurIST toolchain, I gained direct access to the entire functionality of GrAL.
Computational services
The APD concept and its implementation has some important consequences. It means that the same input (APD) can be used to drive completely different simulation packages (a custom preprocessor must be implemented for each package). It also makes long-term storage of computational problems less prone to difficulties with missing backward compatibility. The complete automation of the APD processing makes is the ideal input for computational services (deployed in a service oriented architecture), because it retains its high-level semantics, but does not prescribe any particular solution method. Current service offerings in the field of e.g. computational mechanics are typically tied to a very specific application, and force the client to tailor input to this service more than the problem itself requires. Thus, APDs better support the goal of decoupling services for a specific task from their use.
I used the APD interface to implement service back ends for the simulation applications mentioned before (ANSYS CFX, DC Lattice Boltzmann, FEANOR). In cooperation with Guillem Cantallops and Rodrigo Ruiz (Grid Systems), we set up computational services using the Fura middleware.